-
Notifications
You must be signed in to change notification settings - Fork 34
SmartSeq2 Pipeline Timing and Cost
It could be very complicated to estimate the cost of cloud computing. For google cloud, the billing is on the reserved resource, such as number of cores, size of harddrive and RAM. You will be charged by an hourly rate over the time you use the reserved resource. The pricing also varies between machine type. For example, the hourly rate of using a high specs machine (more nodes and high RAM) is higher than low specs ones. The pricing page lists details.
The cost of processing RNA-Seq on cloud computing depends one several factors. One is running time. Second is how many resources have been reserved during data processing. Third is machine type. It could be tricky to estimate the cost. For example, to run one RSEM job, we can either request min resource, such as 1-core with 4Gb RAM machine or we can request a 4-core and 15Gb machine. The hourly rate of the first type of machine is cheaper than the second one but it takes longer time to finish job.
In this task, we estimated the cost of processing scRNA-Seq data by using google cloud. The testing pipeline included the following modules/steps:
- STAR alignment [request 8-core and 40Gb RAM]
- RSEM to estimate gene counts [request 4-core and 4Gb RAM]
- FeatureCount to calculate gene/exon/transcript counts [request 1-core and 4Gb RAM]
- Picard to collect several sequencing and RNA-Seq specific metrics [request 1-core and 4Gb RAM]
- Python script to parse pipeline output [request 1-core and 2Gb]
Then we tested this pipeline on a published scRNA-Seq dataset, which includes 864 single cells full length RNA-Seq data. And we summarize the total hours(in hour) and total cost(in dollar) in the following table.
| StepName | Program/software | Machine Type | Total Hours | Total Cost | AVG Hours | AVG Cost |
|---|---|---|---|---|---|---|
| Star | Star | n1-highmem-8 | 292.133 | 140.115 | 0.338 | 0.1621 |
| RSEM | RSEM | n1-standard-4 | 151.983 | 29.376 | 0.1759 | 0.034 |
| CollectAlignemntMetrics | Picard | n1-standard-2 | 144.216 | 13.858 | 0.166 | 0.016 |
| CollectRnaMetrics | Picard | n1-standard-2 | 144.555 | 13.890 | 0.167 | 0.0160 |
| CollectInsertSizeMetrics | Picard | n1-standard-2 | 144.116 | 13.849 | 0.1668 | 0.016 |
| CollectDuplicationMetrics | Picard | n1-standard-2 | 144.433 | 13.958 | 0.167 | 0.0161 |
| FeatureCountsUniqueCounts | FeatureCount | n1-standard-2 | 144.266 | 14.179 | 0.167 | 0.0164 |
| FeatureCountsMultiCounts | FeatureCount | n1-standard-2 | 144.483 | 14.200 | 0.1672 | 0.0164 |
| CollectMetricsbySample | python | n1-standard-1 | 144.466 | 7.535 | 0.167 | 0.0087 |
| Total | 1454.65 | 260.9 |
Notice The Total Hours and Total Cost truly reflect the total amount of running one task in Google computing, which not only includes the actually programming runs but also includes the copying/localizing files into/from google cloud VM.
First we calculated the correlation between overall pipeline running time/cost and RNASeq QC metrics. The overall pipeline running time and cost show correlation with several QC metrics, such as PF_BASES,INTERGENIC_BASES and PF_MISMATCH_RATE.
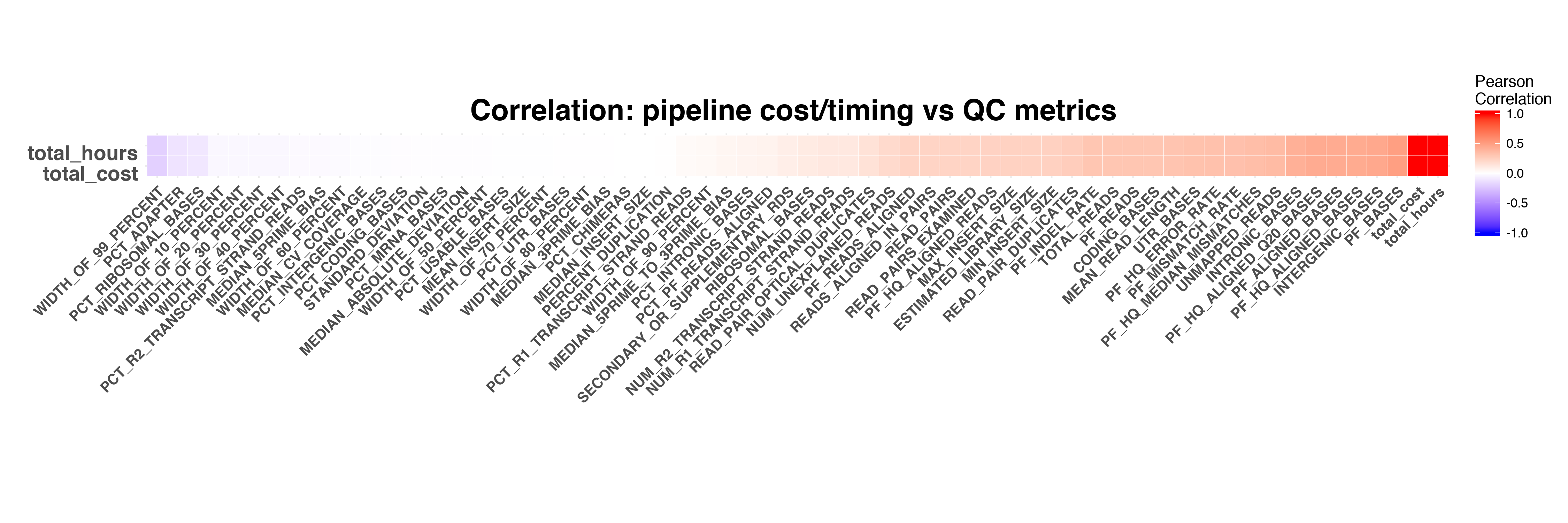
Next we calculated the correlation between each pipeline module's running time&cost and QC metrics
STAR running hours and cost
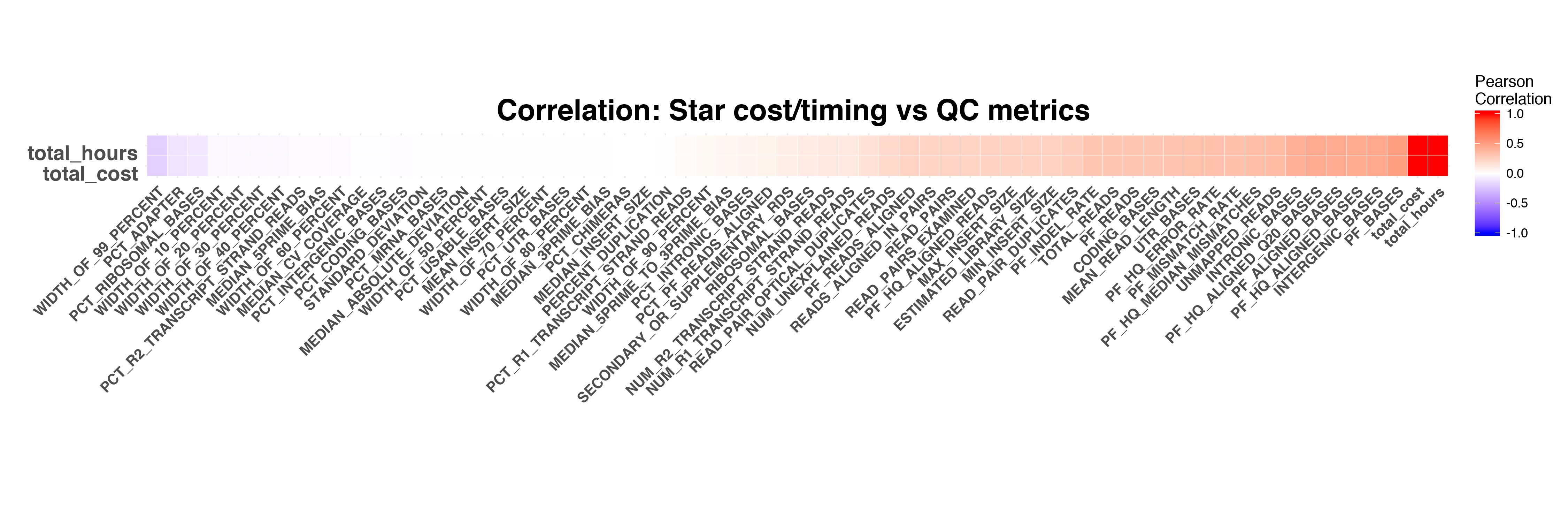 RSEM
RSEM
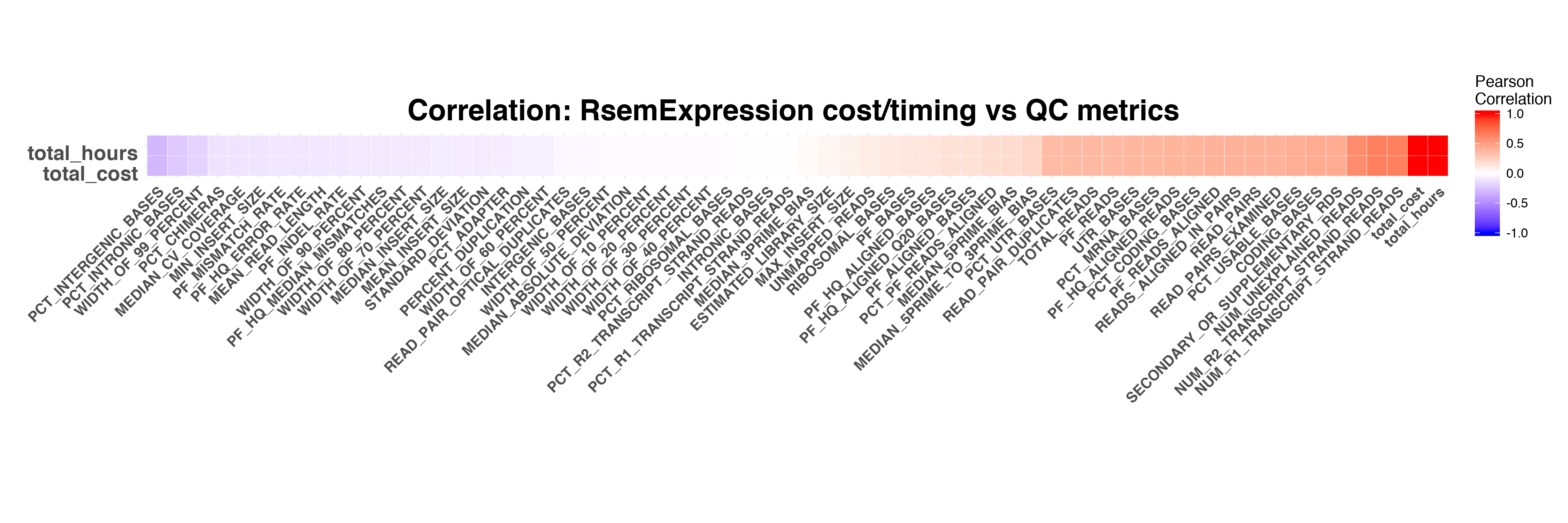 FeatureCounts
FeatureCounts
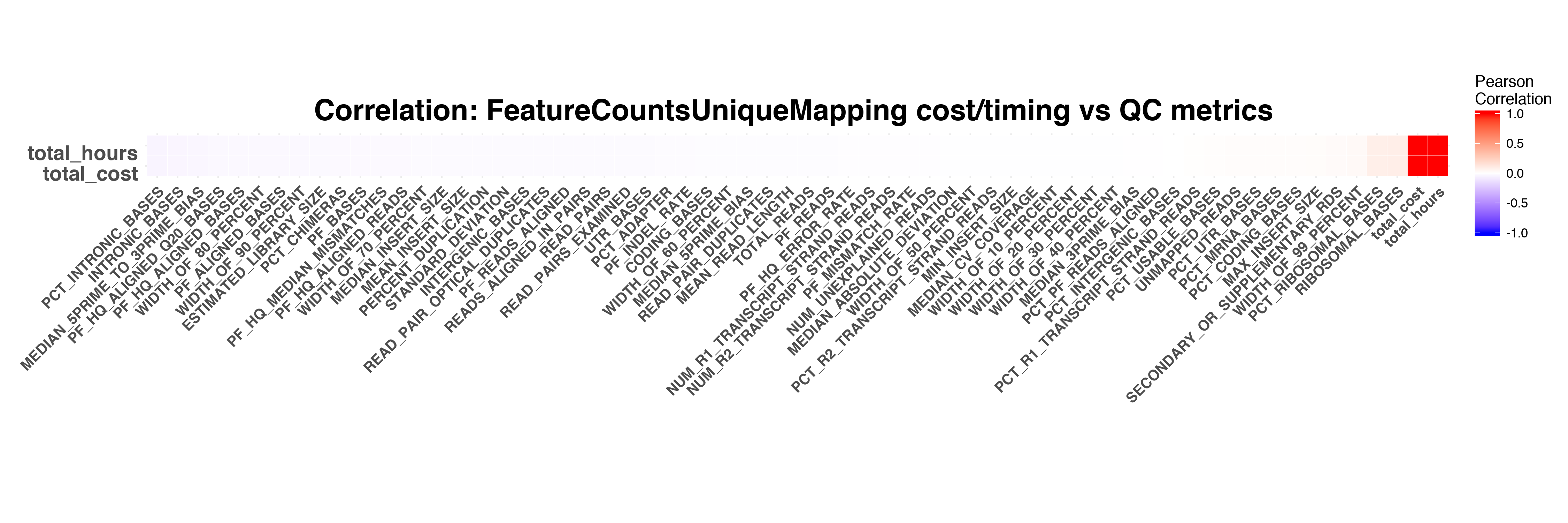 Picard Collect Metrics
Picard Collect Metrics

The timing and cost of STAR and RSEM have stronger correlation to RNASeq QC metrics compared to the timing and cost of Picard and FeatureCounts.
Then we examined the impact of metrics PF_BASES, PF_ALIGNED_BASES,INTRONIC_BASES, PF_MISMATCH_RATE,MEAN_READ_LENGTH on pipeline running time and cost.
In general, PF_BASES,PF_ALIGNED_BASES and INTRONIC_BASES tell us the size of sequencing data and PF_MISMATCH_RATE tell us the quality of alignment.
Impact on STAR
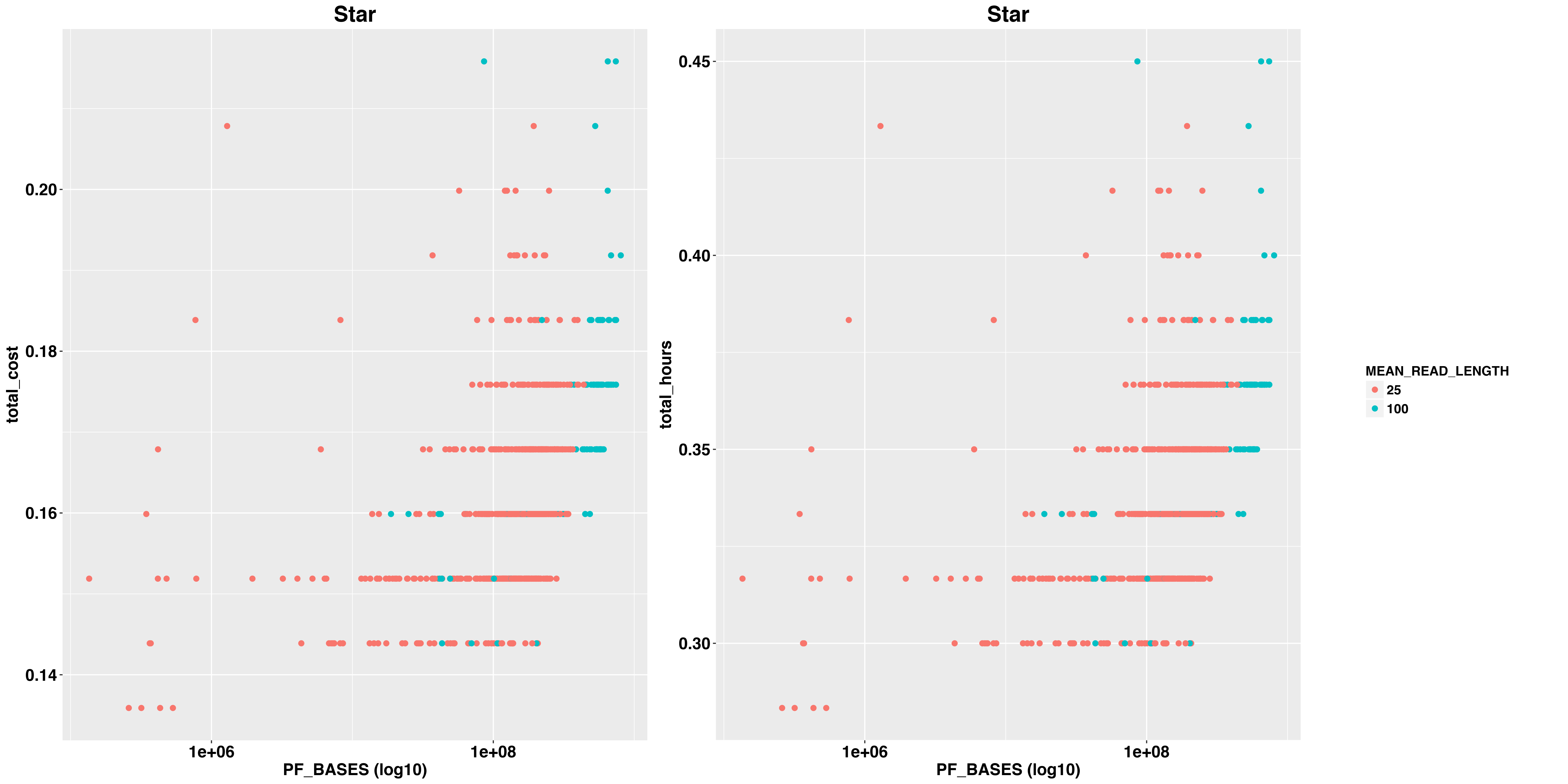




And RSEM
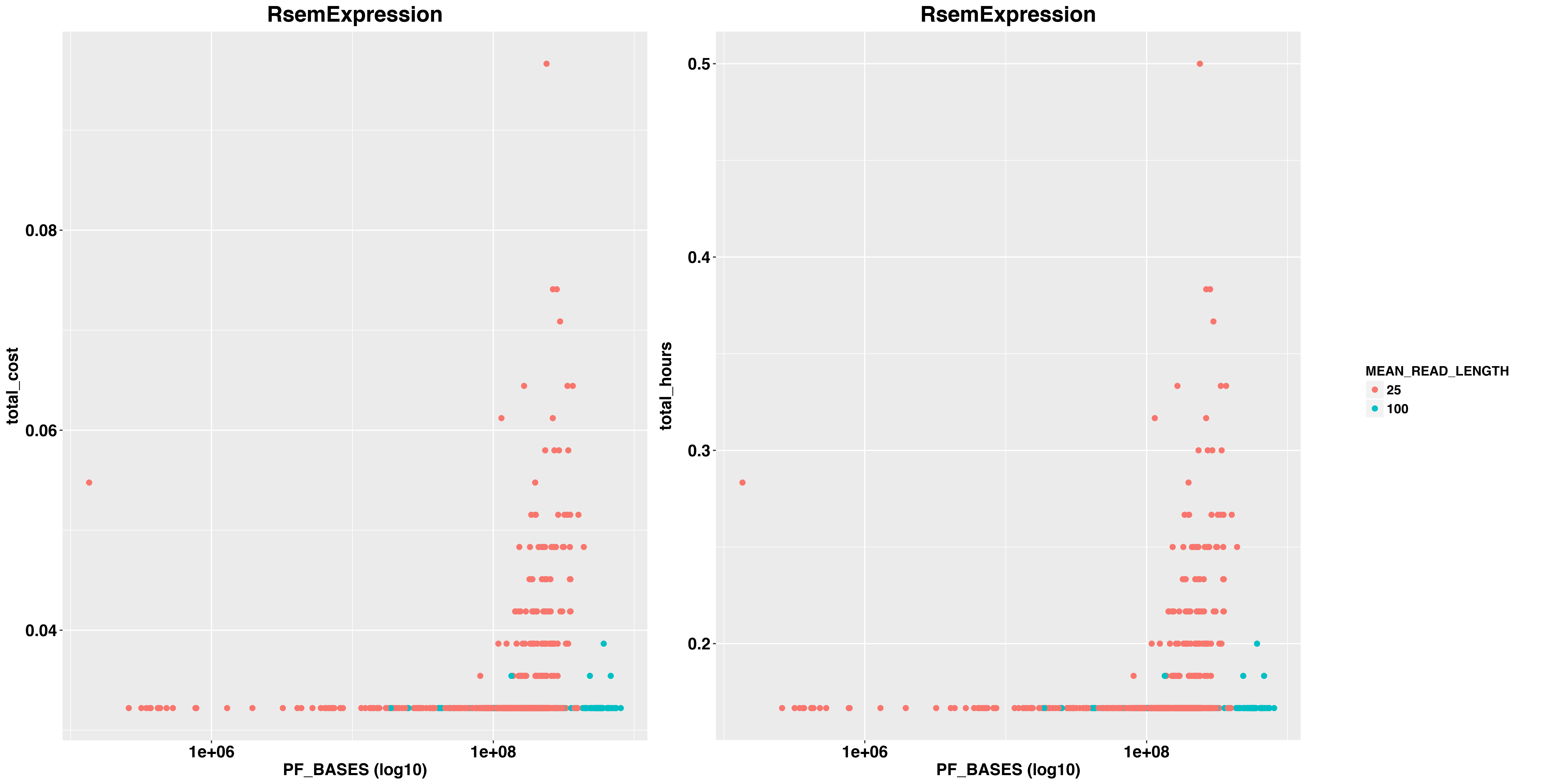


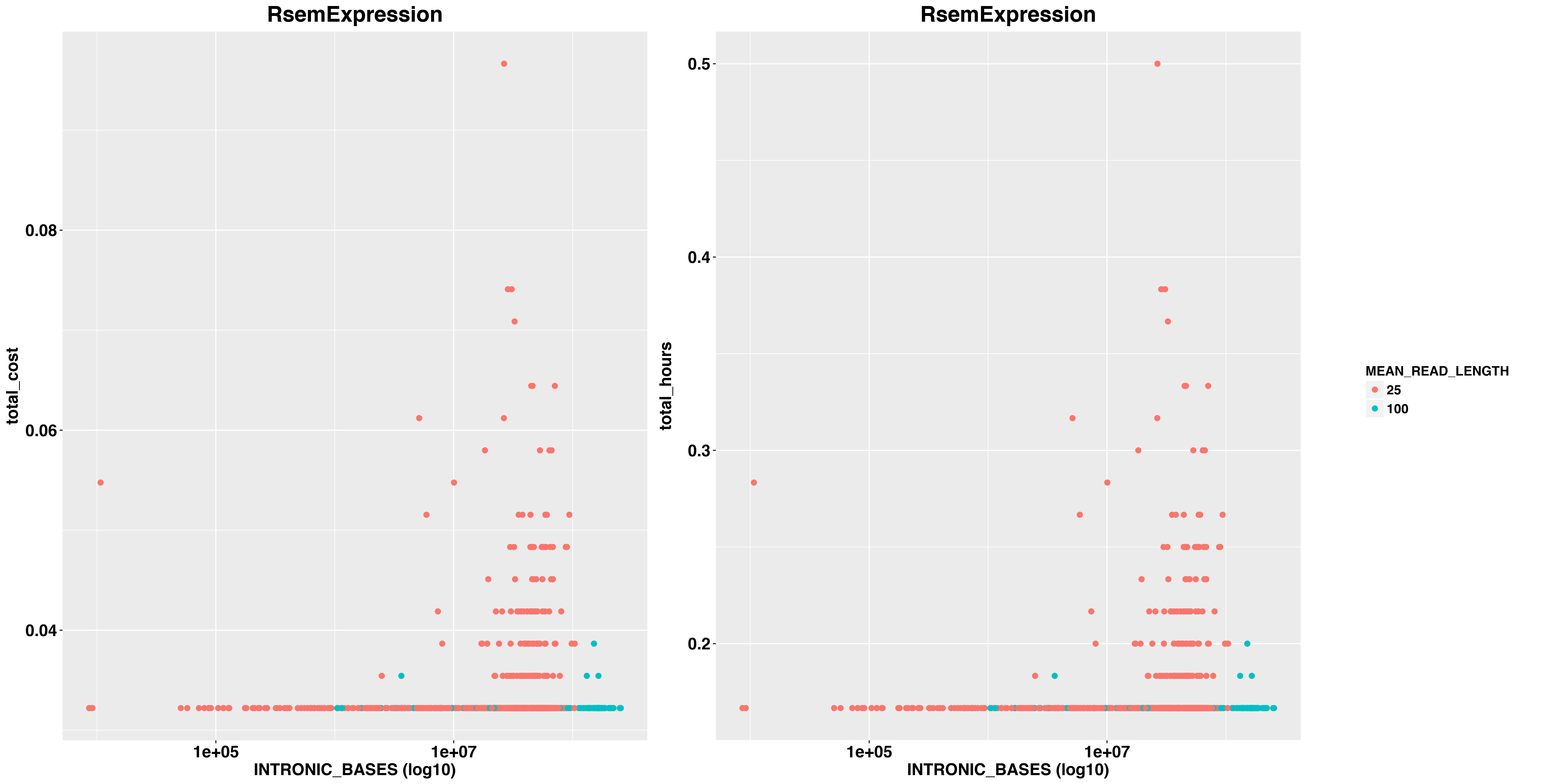 images/costs/total_hours_cost_reads_RsemExpression_BASES_facet_plot.png
images/costs/total_hours_cost_reads_RsemExpression_BASES_facet_plot.png